PySCF UI
PySCF-UI ✨
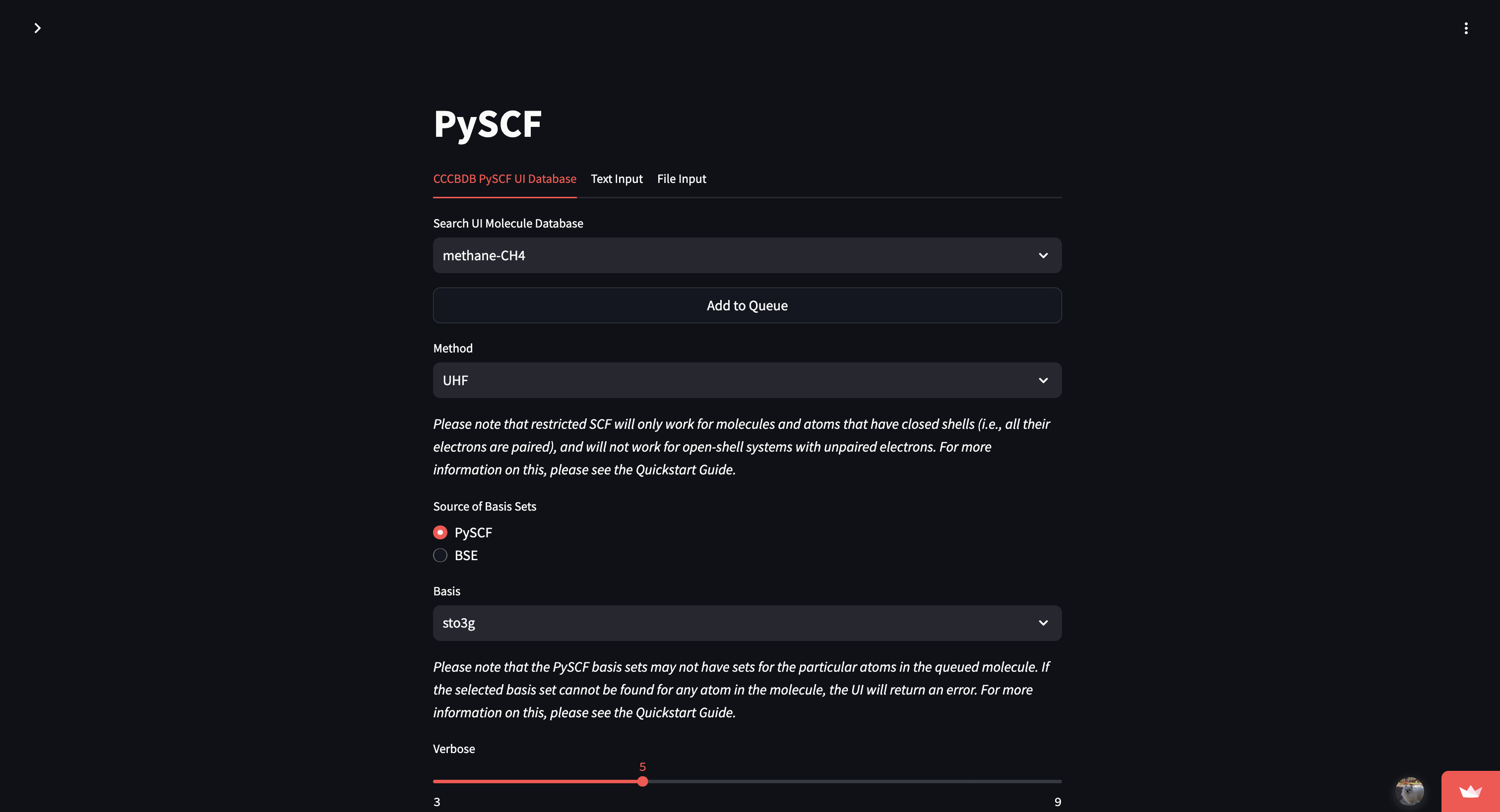
A lightweight Streamlit-based UI for running and inspecting PySCF quantum-chemistry calculations. Designed to make small molecule workflows easy to run, visualize, and compare — perfect for demos, quick experiments, and showcasing to recruiters. 🚀
Key goals: - Provide an approachable web UI for running PySCF calculations. - Collect and display per-molecule runtimes and results. - Include utilities for precomputed molecules and example datasets.
Highlights
- Simple Streamlit app UI for setting up and running calculations.
- Per-molecule runtime tracking and result aggregation (CSV/DataFrame friendly). ⏱️
- RDKit and py3Dmol integration for molecule display and 3D viewing. 🧪🔬
- Precomputed molecules bundled for fast demos.
Quickstart (run locally)
-
Create and activate a virtual environment (recommended):
python -m venv venv source venv/bin/activate # macOS / Linux (zsh) -
Install dependencies:
pip install -r requirements.txt -
Start the Streamlit app:
streamlit run PySCFUI.py
Open the URL printed in the terminal to view the app in your browser.
Notes:
- If your environment or entrypoint differs, main.py or stream.py may be available as alternate entrypoints — check the repository root.
Files of interest
PySCFUI.py— primary Streamlit application (UI + session state). 👀main.py,stream.py— alternate scripts / runners (if present).runner.py— helper routines to execute calculations.precomputed_molecules/— example geometries for quick demos.results/andcomputed/— output folders for results and logs.requirements.txt— Python dependencies used by the project.
Displaying runtimes
The app collects per-molecule runtimes into st.session_state['results'] and you can build a pandas DataFrame from that list. Example snippet to create a DataFrame with one row per molecule:
import pandas as pd
import streamlit as st
rows = []
for rec in st.session_state.get('results', []):
rows.append({
'Molecule': rec.get('Molecule Name'),
'Basis': rec.get('Basis'),
'SCF CPU (s)': rec.get('SCF CPU Runtime'),
'SCF Wall (s)': rec.get('SCF Wall Runtime'),
'Hessian CPU (s)': rec.get('Hessian CPU Runtime'),
'Hessian Wall (s)': rec.get('Hessian Wall Runtime'),
'Real Compute Time (s)': rec.get('Real Compute Time'),
})
df = pd.DataFrame(rows)
st.dataframe(df) # or df.to_html(index=False) + st.markdown(..., unsafe_allow_html=True)If Streamlit raises Arrow serialization errors (e.g. mixed types), force object columns before display:
df = df.astype('object')
st.dataframe(df)To hide the DataFrame index in HTML rendering use df.to_html(index=False) and render via st.markdown(..., unsafe_allow_html=True).
Development notes / legacy entrypoints
- If you prefer or need the older entrypoint, you can run the legacy Streamlit script directly:
# start the older entrypoint
streamlit run stream.py-
test.pyexists as a lightweight harness to exercise functions and pipeline pieces without launching the full Streamlit UI. Use it for quick, iterative checks (for example:python test.py). -
Precomputed molecule files belong in the
precomputed_molecules/folder. Requirements:- Filename must end with
.geom.txt(e.g.methane-CH4.geom.txt). - Follow the same formatting as the files already in the folder: plain-text geometry lines (atom label and XYZ coordinates) and the same header/footer conventions used by the repo. Copy an existing file as a template when adding new entries.
- Filename must end with
Full Steps (quick recap)
- make python environment (if not already created):
python -m venv venv - activate environment:
source venv/bin/activate - install dependencies (if not already installed):
pip install -r requirements.txt - run app (primary entrypoint):
streamlit run PySCFUI.py - or run legacy entrypoint:
streamlit run stream.py - open link printed in terminal in web browser and use it!
Dependencies (see requirements.txt)
- main ones used by the project:
- pyscf
- rdkit
- streamlit
- altair
- pandas
- numpy
- matplotlib
- py3Dmol